- 2026年2月11日
- 2026年2月11日
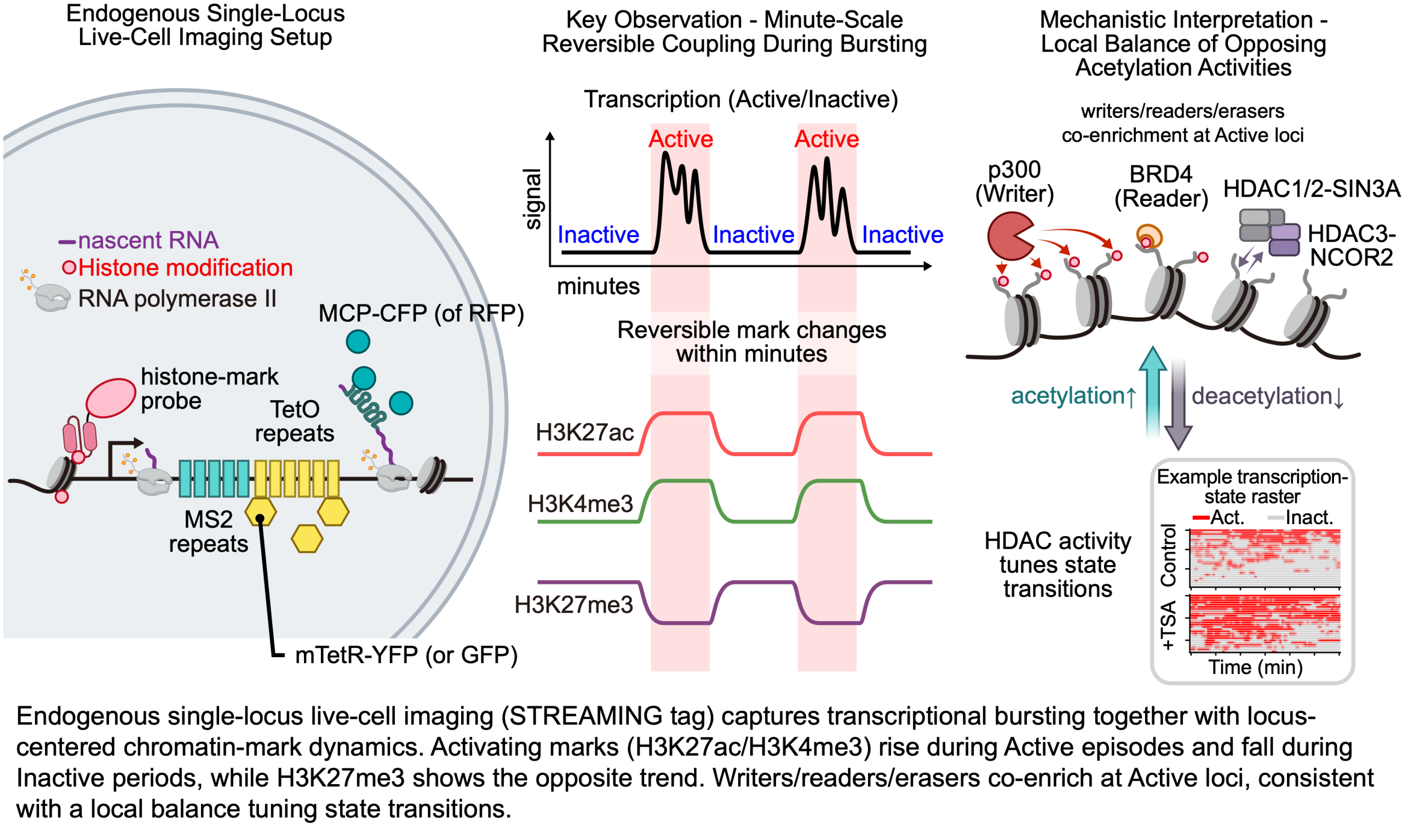
New bioRxiv preprint: Minute-scale coupling of chromatin marks and transcriptional bursts
We have posted a new preprint on bioRxiv: “Minute-scale coupling of chromatin marks and transcriptional bursts” (https://www.biorxiv.org/cgi/content/short/2026.02.08.704500v1). In this study, we combine single-gene live-cell imaging at endogenous loci with fluorescent probes for histone modifications (mintbody/ChromID) to capture transcriptional bursting together with locus-centered chromatin-mark dynamics in mouse embryonic stem cells. We find that activating chromatin marks (H3K27ac/H3K4me3) increase during transcriptionally Active episodes and decrease during Inactive episodes, whereas a Polycomb-associated repressive mark (H3K27me3) shows the opposite tendency. We further show that acetylation-related regulators, including selected deacetylase modules, co-accumulate at Active loci, and that HDAC inhibition reshapes Active/Inactive dwell times, supporting a model in which opposing enzymatic activities tune state transitions on minute timescales. This work was led by graduate students Xiohui Gao and Chaebeen Ko, and was conducted in collaboration with Prof. Yasuyuki Ohkawa lab and Prof. Akihito Harada lab (Kyushu University), and Prof. Hiroshi Kimura (Tokyo Institute of Science). We […]